Discuss the estimation of the number of transcriptional enhancers in the human genome, and how to study their regulatory activity
Introduction
An enhancer is a DNA region that can be bound with transcription factor protein or activators to up regulate transcription of a gene or genes (Pennacchio et al., 2013). Transcription of a gene refers to the start of gene expression when a particular DNA segment is transcribed into RNA using RNA polymerase. Enhancers are cis-acting and the human genome contains many of those regions. Enhancers belong to a class of transcriptional regulatory elements that includes promoters, silencers, insulators, and locus control regions. Enhancer's primary role is regulating the process of gene expression. Enhancers are hard to locate in the human genome as they are widely distributed (Pennacchio et al., 2013). They also contain poorly understood sequence features. Wide distribution and inability to understand enhancer region fully, leads to the difficulty in locating and computing the number of transcriptional enhancers (Pennacchio et al.,
2013).
Location of Transcriptional Enhancers
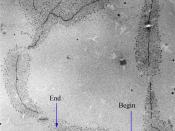
Enhancers are located either upstream or downstream of the gene it regulates. It is not necessary for the enhancer to be near the transcription initiation site. Studies have found that some enhancers are located several thousand bases upstream or downstream from transcription initiation site (Pennacchio et al., 2013). Enhancers are bound by activator proteins/transcription factors in a cis-acting manner, and hence do not act in the promoter region (Shlyueva, Stampfel and Stark, 2014). Each octamer has about 147 base pair. Active enhancer and promoters can be identified by checking for depletion of nucleosome, some nucleosomes that flank active enhancers show specific histone modifications as shown in figure 1.
a) Shows Intricate regulatory functions of the different enhancer (A and B), (4* 1 Histone core) (588 base pair) downstream...