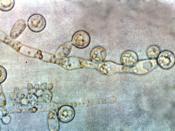
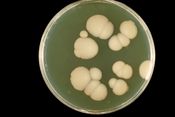
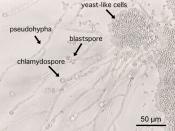
With reference to named examples, discuss the use of genomics in understanding the pathogenesis and detection of human fungal infections.
1 Introduction
1.1 History of fungal genomics:
The thallophytic plant offshoot, the fungus is known to be one of the pathogenic species that cause infection in humans. Progress on fungal genomes has been particularly limited at present, with only recent developments made for instance in 1996 the work published on the genome of Saccharomyces cerevisiae (Goffeau et al. 1996).
As a result the dawn of a new genre in human disease has blossomed. Related by Lee et al (Lee et al. 2002), this supposedly was the first dramatic large-scale study on the eukaryotic fungal diseases. This warming investigation led the way to genomics as now many researchers have mimicked Goffeau et al and have plowed out genes of fungi and their functions in pathogenesis, and human fungal infections (Jorgensen et al.
2002).
Scientists working on human genes have now a broader understanding in the connection of gene function and pathogenesis, as there is many bodies geared into action and have advanced in fungal research. Once such include MIPS - Munich information center for protein sequences. Where their services include online genome databases that show the study of all of the nucleotide sequences, including structural genes of fungal genomes (Andreoli C et al 2004 - from MIPS website).
Genome sequence from these established fungi will aid our understanding of the eukaryotic proteome and ultimately help to detect and manipulate fungi for our own purposes, especially for treatment from the development of disease. Nonetheless a growing concern is on the lack of enhancement in this area as fungal genome sequence can provide the keys to underlining the basis of fungal interaction and human medicine.
1.2 Introduction to Fungi:
It roared itself...