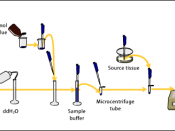
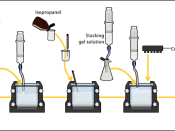
AimsTo analyze four protein samples (Crude extract, AS1, AS2, AS3) using native (non-denaturing) polyacrylamide gel electrophoresis (PAGE).
To identify a number of proteins and isoenzyme bands by staining with Coomassie Blue and isoenzyme staining and to determine their relative mobilities.
IntroductionThe resolution of separate components in a protein mixture is obtained most clearly using an electrophoretic method. It is used to characterize a purified protein preparation and requires only a small amount (5 to 25õg) of protein. In this practical, native polyacrylamide gel is used. This involves running the sample in a buffer at a pH suitable for protein stability and to remain in their native form.
The pH of the resolving gel is 8.8 and is alkaline to keep most proteins negatively charged to enable movement towards the anode. A 10% resolving gel is used so the pore size of the gel enables medium sized proteins to pass through.
A higher percentage of acrylamide/bis solution would give smaller pores. A discontinuous buffer system was used. The stacking gel allows protein to be concentrated before they enter the resolving gel, increasing resolution by forming a sharp band of protein.
Coomassie Blue binds with a large range of proteins almost stoichiometrically and isoenzyme staining detects enzyme activity in a coupled manner. The comparison of the stainings enables estimation of the mass to charge ratio of ñ-esterase isoenzymes in relation to the other proteins in the extract. Isoenzyme staining is based on the formation of an insoluble dye or precipitate final product (produced with the mixture of solution A and solution B) upon interaction with the enzyme of interest. Isoenzyme staining is used as an indication of particular enzyme activity. Thus, enzymes must remain functional and only native gel electrophoresis can be carried out. Even though sodium dodecylsulphate-PAGE (SDS-PAGE) can be calibrated...