Genetic Fingerprinting.
It has been found in research that sections of DNA which do not code for part of a gene contains highly repetitive sequences of bases. These are called Variable Number Tandem Repeats, {VNTR's}. A genetic fingerprint is a method of revealing the differences in size of VNTR's between individuals. Firstly a DNA sample is extracted from the cell. Restriction enzymes then cut sections of the DNA close to but not within the VNTR regions. The DNA fragments produced are then separated according to size using aragose gel electrophoresis. The smaller the fragments, the further they travel to the anode. The fragments produced are invisible at this stage and so the fragments are transferred to a nylon membrane using southern blotting which involves adding a layer of absorbent paper to the nylon membrane. The DNA is then drawn upwards by capillary motion. The DNA fragments are then denatured by heating to give single stranded DNA. A radioactive isotope of phosphorus with a base sequence identical to one of the VNTR sequences is used to locate the particular bands. This is also known as a DNA probe and it binds to its complementary single stranded DNA. Any excess probe is washed off. The probes are accurately located by placing X-ray film over the nylon membrane.




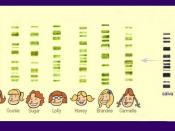
Short.
Descriptive but no explanation. A longer essay with Explanation would be helpful. The organiztion could use some attention no paragarphs or closeing statements.
2 out of 2 people found this comment useful.